A table of the RNA codons together with the amino acids they encode.
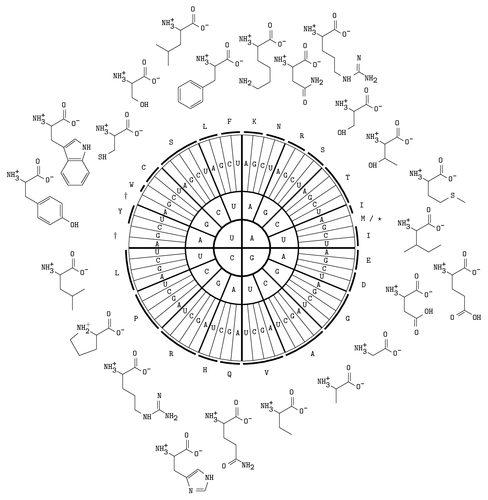
Edit and compile if you like:
% RNA codons
% Author: Florian Hollandt
\documentclass{article}
\usepackage{tikz}
\usepackage[active,tightpage]{preview}
\PreviewEnvironment{tikzpicture}
\setlength\PreviewBorder{5pt}%
\setlength\oddsidemargin{0in}
\begin{document}
\ttfamily
\footnotesize
\pagestyle{empty}
\begin{center}
\begin{tikzpicture}
\tikzstyle{every node}=[inner sep=1.7pt,anchor=center]
% to_x and from_x styles denote bonds terminating or starting in labeled nodes. x denotes the number of letters in the node label.
\tikzstyle{to_1}=[shorten >=5pt]
\tikzstyle{to_1i}=[shorten >=6pt]
\tikzstyle{to_2}=[shorten >=7pt]
\tikzstyle{to_3}=[shorten >=8pt]
\tikzstyle{from_1}=[shorten <=5pt]
\tikzstyle{from_1i}=[shorten <=6pt]
\tikzstyle{from_2}=[shorten <=8pt]
\begin{scope}
\draw [ultra thick] circle(1cm);
\draw [ultra thick] (0:4)--(180:4) (90:4)--(270:4);
\foreach \a/\l in {45/A,135/G,225/C,315/U}{
\node at (90-\a:0.5cm) {\l};
}
\draw [very thick] circle(2cm);
\foreach \A in {90,0,270,180}{
\foreach \a/\l in {22.5/A,45/G,67.5/C,90/U}{
\draw [very thick] (\A+\a:1) -- (\A+\a:4);
\node at (\A-\a+11.25:1.5) {\l};
}
}
\draw circle(4cm) (0:4)--(180:4) (90:4)--(270:4);
\foreach \A in {90,180,270,0}{
\foreach \a in {0,22.5,45,67.5}{
\foreach \i/\l in {5.625/A,11.25/G,16.875/C,22.5/U}{
\draw (\A+\a+\i:2) -- (\A+\a+\i:4);
\node at (\A-\a-\i+2.8125:3) {\l};
}
}
}
\end{scope}
\begin{scope}[scale=0.5] % Lysine
\draw[ultra thick,shorten >=2pt,shorten <=2pt] (90:8.2)
arc(90:90-2*5.625:8.2);
\path (90-0.8*5.625:14.3) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw[to_3] (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node (Cc) {}
-- ++(270:1) node (Cd) {}
-- ++(210:1) node (Ce) {}
-- ++(150:1) node (Cf) {NH$_{\mbox{2}}$};
\end{scope}
\begin{scope}[scale=0.5] % Asparagine
\draw[ultra thick,shorten >=2pt,shorten <=2pt] (90-2*5.625:8.2)
arc(90-2*5.625:90-4*5.625:8.2);
\path (90-3.5*3.625-3:13.3) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw[to_2] (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node (Cc) {}
-- +(30:1) node (Cd) {NH$_{\mbox{2}}$};
\draw[to_1i] (Cc.center) -- +(270:1) node (O) {};
\draw[to_1] (Cc.210) -- (O.150);
\path (O.center) node {O};
\end{scope}
\begin{scope}[scale=0.5] % Arginine
\draw[ultra thick,shorten >=2pt,shorten <=2pt] (90-22.5:8.2)
arc(90-22.5:90-33.75:8.2);
\path (90-3.7*5.625:16) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw[to_2] (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node (Cc) {}
-- ++(270:1) node (Cd) {}
-- ++(330:1) node (NH1) {NH};
\draw[from_2,to_3] (NH1.center) -- ++(30:1) node (Ce) {}
-- ++(330:1) node {NH$_{\mbox{2}}$};
\draw[to_1i] (Ce.center) -- ++(90:1) node (N2) {};
\draw[to_1] (Ce.150) -- (N2.210);
\path (N2) node {N};
\end{scope}
\begin{scope}[scale=0.5] % Serine
\draw[ultra thick,shorten >=1pt,shorten <=2pt] (90-22.5-2*5.625:8.2)
arc(90-33.75:90-33.75-11.25:8.2);
\path (90-7*5.625:12.5) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw[to_2] (zero.center) -- ++(270:1) node(Cb){} -- ++(210:1) node (Cc) {OH};
\end{scope}
\begin{scope}[scale=0.5] % Threonine
\draw[ultra thick,shorten >=1pt,shorten <=2pt] (90-45:8.2)
arc(90-45:90-67.5:8.2);
\path (90-45-0.8*11.25:12.5) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw[to_2] (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node (Cc) {} (Cb.center)
-- +(210:1) node {OH};
\end{scope}
\begin{scope}[scale=0.5] % Methionine
\draw[ultra thick,shorten >=1pt,shorten <=2pt] (90-67.5:8.2)
arc(90-67.5:90-67.5-5.625:8.2);
\path (90-67.5-0.5*5.625:14) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw[to_1] (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node (Cc) {}
-- ++(30:1) node (Cd) {S};
\draw[from_1] (Cd.center) -- +(330:1);
\end{scope}
\begin{scope}[scale=0.5] % Isoleucine
\draw[ultra thick,shorten >=1pt,shorten <=2pt] (0:8.2)
arc(0:11.25:8.2);
\path (1.0*5.625:12.4) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node (Cc) {}
-- +(30:1) node (Cd) {} (Cb.center)
-- +(210:1) node (Ce) {};
\end{scope}
\begin{scope}[scale=0.5] % Glutamic acid
\draw[ultra thick,shorten >=1pt,shorten <=2pt] (0:8.2)
arc(0:-11.25:8.2);
\path (-1.3*5.625:15) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw[to_1i] (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node (Cc) {}
-- ++(270:1) node (Cd) {}
-- ++(330:1) node (NH) {OH};
\draw[to_1] (Cd.center) -- +(210:1) node (O) {};
\draw[to_1i] (Cd.270) -- (O.300);
\path (O.center) node {O};
\end{scope}
\begin{scope}[scale=0.5] % Aspartic acid
\draw[ultra thick,shorten >=1pt,shorten <=2pt] (-11.25:8.2)
arc(-11.25:-22.5:8.2);
\path (-11.25-5.625:12) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw[to_2] (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node (Cc) {}
-- +(30:1) node (Cd) {OH};
\draw[to_1i] (Cc.center) -- +(270:1) node (O) {};
\draw[to_1] (Cc.210) -- (O.150);
\path (O.center) node {O};
\end{scope}
\begin{scope}[scale=0.5] % Glycine
\draw[ultra thick,shorten >=1pt,shorten <=2pt] (-22.5:8.2)
arc(-22.5:-45:8.2);
\path (-33.75-1*5.625:12) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\end{scope}
\begin{scope}[scale=0.5] % Alanine
\draw[ultra thick,shorten >=1pt,shorten <=2pt] (-45:8.2)
arc(-45:-68.25:8.2);
\path (-45-11.25:12) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw (zero.center) -- ++(270:1) node(Cb){};
\end{scope}
\begin{scope}[scale=0.5] % Valine
\draw[ultra thick,shorten >=1pt,shorten <=2pt] (-68.25:8.2)
arc(-68.25:-90:8.2);
\path (-68.25-0.8*11.25:12) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node (Cc) {};
\end{scope}
\begin{scope}[scale=0.5] % Glutamine
\draw[ultra thick,shorten >=1pt,shorten <=2pt] (-90:8.2)
arc(-90:-101.25:8.2);
\path (-90.25-5.625:12.5) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_2] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw[to_3] (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node (Cc) {}
-- ++(270:1) node (Cd) {}
-- ++(330:1) node (NH) {NH$_{\mbox{2}}$};
\draw[to_1] (Cd.center) -- +(210:1) node (O) {};
\draw[to_1i] (Cd.270) -- (O.300);
\path (O.center) node {O};
\end{scope}
\begin{scope}[scale=0.5] % Histidine
\draw[ultra thick,shorten >=1pt,shorten <=2pt] (-101.25:8.2)
arc(-101.25:-101.25-11.25:8.2);
\path (-101.25-1.2*5.625:15.5) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node(Cc){};
\draw[to_2] (Cc.center) -- ++(108-1*72:1) node (Cd) {}
-- ++(108-2*72:1) node (Ce) {NH};
\draw[from_1,to_1] (Ce.center) -- ++(108-3*72:1) node (Cf) {}
-- ++(108-4*72:1) node (Cg) {};
\draw[from_1] (Cg.center) -- (Cc.center);
\draw (Cc.198+2*72) -- (Cd.198+1*72);
\draw[from_1] (Cg.72) -- (Cf.198+4*72);
\draw (Cg.center) node {N};
\end{scope}
\begin{scope}[scale=0.5] % Arginine
\draw[ultra thick,shorten >=2pt,shorten <=2pt] (-90-22.5:8.2)
arc(-90-22.5:-90-45:8.2);
\path (-90-7.7*5.625:12.3) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw[to_2] (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node (Cc) {}
-- ++(270:1) node (Cd) {}
-- ++(330:1) node (NH1) {NH};
\draw[from_1i,to_3] (NH1.center)-- ++(30:1) node (Ce) {}
-- ++(330:1) node {NH$_{\mbox{2}}$};
\draw[to_1] (Ce.center) -- ++(90:1) node (N2) {};
\draw[shorten >=4pt] (Ce.150) -- (N2.210);
\path (N2) node {N};
\end{scope}
\begin{scope}[scale=0.5] % Proline
\draw[ultra thick,shorten >=2pt,shorten <=2pt] (-90-45:8.2)
arc(-90-45:-90-45-22.25:8.2);
\path (-90-10.5*5.625:12) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_2] (zero.center) -- ++(150:1) node (nh) {NH$_{\mbox{2}}^+$};
\draw (zero.center) -- ++(270:1) node(Cb){};
\path (Cb.center) -- +(150:1) node (x) {};
\path (x.center) +(170:1) node (Cd) {};
\path (x.center) +(250:1) node (Cc) {};
\draw[to_3] (Cb.center) -- (Cc.center)
-- (Cd.center)
-- (nh.center);
\end{scope}
\begin{scope}[scale=0.5] % Leucine
\draw[ultra thick,shorten >=2pt,shorten <=2pt] (180:8.2)
arc(180:180+22.25:8.2);
\path (-90-14.5*5.625:13) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node (Cc) {}
-- +(30:1) node (Cd) {} (Cc.center)
-- +(270:1) node (Ce) {};
\end{scope}
\begin{scope}[scale=0.5] % Tyrosine
\draw[ultra thick,shorten >=2pt,shorten <=2pt] (180-11.25:8.2)
arc(180-11.25:180-22.5:8.2);
\path (180-3*5.625:16) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw (zero.center) -- ++(270:1) node(Cb){};
\draw (Cb.center) -- ++(330:1) node (Cc) {}
-- ++(30:1) node (Cd) {}
-- ++(330:1) node (Ce) {}
-- ++(270:1) node (Cf) {}
-- ++(210:1) node (Cg) {}
-- ++(150:1) node (Ch) {}
-- ++(90:1);
\draw (Cc.330) -- (Cd.270);
\draw (Ce.210) -- (Cf.150);
\draw (Cg.90) -- (Ch.30);
\draw[to_1i] (Cf.center) -- +(330:1) node (OH) {OH};
\end{scope}
\begin{scope}[scale=0.5] % Tryptophane
\draw[ultra thick,shorten >=2pt,shorten <=2pt] (180-22.5-5.625:8.2)
arc(180-22.5-5.625:180-22.5-11.25:8.2);
\path (180-22.5-1.8*5.625:16) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node(Cc){};
\draw[to_2] (Cc.center) -- ++(108-1*72:1) node (Cd) {}
-- ++(108-2*72:1) node (Ce) {NH};
\draw[from_1](Ce.center) -- ++(108-3*72:1) node (Cf) {}
-- ++(108-4*72:1) node (Cg) {};
\draw (Cg.center) -- (Cc.center);
\draw (Cc.198+2*72) -- (Cd.198+1*72);
\draw (Cg.72) -- (Cf.198+4*72);
\draw (Cg.center) -- ++(240:1) node (Ch) {}
-- ++(300:1) node (Ci) {}
-- ++(0:1) node (Cj) {}
-- ++(60:1) node (Ck) {}
-- ++(120:1) node (Cl) {};
\draw (Ch.0) -- (Ci.60);
\draw (Cj.120) -- (Ck.180);
\end{scope}
\begin{scope}[scale=0.5] % Cysteine
\draw[ultra thick,shorten >=2pt,shorten <=2pt] (180-45+11.25:8.2)
arc(180-45+11.25:180-45:8.2);
\path (180-45+11.25-1*7.625:12) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw[to_2] (zero.center) -- ++(270:1) node(Cb){}
-- ++(210:1) node (Cc) {SH};
\end{scope}
\begin{scope}[scale=0.5] % Serine
\draw[ultra thick,shorten >=1pt,shorten <=2pt] (90+45:8.2)
arc(90+45:90+45-22.5:8.2);
\path (90+45-11.25+0*5.625:14) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw[to_2] (zero.center) -- ++(270:1) node(Cb){}
-- ++(330:1) node (Cc) {OH};
\end{scope}
\begin{scope}[scale=0.5] % Leucine
\draw[ultra thick,shorten >=2pt,shorten <=2pt] (90+22.5:8.2)
arc(90+22.5:90+11.25:8.2);
\path (90+22.5-1.2*5.625:16) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw (zero.center) -- ++(270:1) node(Cb){}
-- ++(210:1) node (Cc) {}
-- +(150:1) node (Cd) {} (Cc.center)
-- +(270:1) node (Ce) {};
\end{scope}
\begin{scope}[scale=0.5] % Phenylalanine
\draw[ultra thick,shorten >=2pt,shorten <=2pt] (90+11.25:8.2)
arc(90+11.25:90:8.2);
\path (90+1.5*5.625:13) node (zero) {};
\draw[to_2] (zero.center) -- ++(30:1) node (CO) {}
-- +(330:1) node [anchor=base] {O$^{\mbox{-}}$};
\draw[to_1] (CO.center) -- +(90:1) node (Od) {O};
\draw[to_1i] (CO.30) -- +(90:1);
\draw[to_3] (zero.center) -- ++(150:1) node {NH$_{\mbox{3}}^{\mbox{+}}$};
\draw (zero.center) -- ++(270:1) node(Cb){};
\draw (Cb.center) -- ++(210:1) node (Cc) {}
-- ++(150:1) node (Cd) {}
-- ++(210:1) node (Ce) {}
-- ++(270:1) node (Cf) {}
-- ++(330:1) node (Cg) {}
-- ++(30:1) node (Ch) {}
-- ++(90:1);
\draw (Cc.210) -- (Cd.270);
\draw (Ce.330) -- (Cf.30);
\draw (Cg.90) -- (Ch.150);
\end{scope}
\node at (90-1*5.625:4.5) {K};
\node at (90-3*5.625:4.5) {N};
\node at (90-5*5.625:4.5) {R};
\node at (90-7*5.625:4.5) {S};
\node at (90-10*5.625:4.5) {T};
\node at (90-12.5*5.625:4.5) {I};
\node at (90-13.7*5.625:4.7) {M / $\star$};
\node at (90-15*5.625:4.5) {I};
\node at (90-17*5.625:4.5) {E};
\node at (90-19*5.625:4.5) {D};
\node at (90-22*5.625:4.5) {G};
\node at (90-26*5.625:4.5) {A};
\node at (90-30*5.625:4.5) {V};
\node at (90-33*5.625:4.5) {Q};
\node at (90-35*5.625:4.5) {H};
\node at (90-38*5.625:4.5) {R};
\node at (90-42*5.625:4.5) {P};
\node at (90-46*5.625:4.5) {L};
\node at (90-49*5.625:4.5) {$\dagger$};
\node at (90-51*5.625:4.5) {Y};
\node at (90-52.3*5.625:4.5) {$\dagger$};
\node at (90-53.3*5.625:4.5) {W};
\node at (90-55*5.625:4.5) {C};
\node at (90-58*5.625:4.5) {S};
\node at (90-61*5.625:4.5) {L};
\node at (90-63*5.625:4.5) {F};
\end{tikzpicture}
\end{center}
\end{document}
Click to download: rna-codons-table.tex • rna-codons-table.pdf
Open in Overleaf: rna-codons-table.tex


